Journal of Plant Biochemistry & Physiology
Open Access
ISSN: 2329-9029
ISSN: 2329-9029
Research Article - (2019)Volume 7, Issue 3
The present investigation was implemented to search for high efficient bacterial strains in bio-remediating the toxic influence of Lead chloride (Pb Cl2) on faba bean plants grown in sandy soil supplemented with various concentrations of Pb Cl2. Five strains were isolated from Al-Taif, Al-Baha and Al-Madinah Al-Munawrah governorates from the rhizosphere of cucumber, fig and tomato plants respectively, KSA. Regarding efficiency of these isolated strains in remediating PbCl2 in the nutrient broth medium, the absorbed quantities ranged from 189 to 417 mg/l while the whole amount recorded in control was 612 mg/l. The isolated bacterial strains were genetically identified as Bacillus amyloliquefaciens subsp. plantarum Trigo Core 1448, Bacillus amyloliquefaciens subsp. plantarum UCMB 5033, Bacillus subtilis strain CCM 1999, Serratia marcescens strain WW4 and Bacillus amyloliquefaciens CC178. Bacillus amyloliquefaciens subsp. plantarum UCMB 5033 revealed the highest efficiency in absorbing Pb that reached 417 mg/l against 612 mg/l for the controls. Bioremediation of Pb-polluted soil with Bacillus amyloliquefaciens significantly increased faba bean plant height, plant fresh weight, dry weight, plant N, P and K concentrations compared to the non-inoculated treatments.
Bio-remediation; Lead-tolerant bacterial strains
Lead is a naturally occurring, bluish-gray metal usually found as a mineral combined with other elements, such as sulphur (i.e., PbS, PbSO4), or oxygen (PbCO3), and ranges from 10 to 30 mg/kg in the earth’s crust [1]. Typical mean Pb concentrations for surface soils worldwide ranges from 10 to 67 mg/kg [2]. Lead enters the environment from industrial activities, such as production of batteries, pigments, metal smelting and manufacture of lead arsenate insecticides or through natural processes like soil erosion, volcanic emission [3]. The total lead concentration in industrial area can reach up to 10000 mg kg-1, and the average value in soil may reach from 10 to 100 mg kg-1. Lead is a major pollutant that is found in soil, water and air, and is hazardous waste and highly toxic to human, animals, plants and microbes [4]. The risk of lead poisoning through the food chain rises as the soil lead level rises above the concentration of 300 ppm [5]. Bioremediation using microorganisms and/or plants is used to free soils from lead contamination, and Blaylock et al. [6] reported 50% to 60% saving when bioremediation was used for the treatment of 1 acre of Pb polluted soil compared to the non-remediated treatment. Bioremediation processes are very attractive in comparison with physicochemical methods such as electrochemical treatment, ion exchange, precipitation, reverse osmosis, evaporation, and sorption for heavy metal removal techniques because they can have lower cost and higher efficiency at low metal concentrations [7,8]. Various lead resistant mechanisms employed by lead resistant bacteria includes efflux mechanism, extracellular sequestration, biosorption, precipitation, alteration in cell morphology, enhanced siderophore production and intracellular lead bioaccumulation [9].
Several microorganisms, especially bacteria (Bacillus subtilis, Pseudomonas putida, and Enterobacter cloacae) have been successfully used for the bioremediation of heavy metals [10-12] isolated several micro-organisms resistant to lead (Pb) among them is the gram-positive bacteria Bacillus cereus, Arthrobacter sp., and Corynebacterium sp. And the gram-negative bacteria Pseudomonas marginalis, P. vesicularis and Enterobacter sp., and fungi Saccharomyces cereviside and Penicillium sp. Six bacterial strains were found to be most resistant isolates and amongst them one isolate showed maximum resistance to lead which was identified as Bacillus cereus, Scanning Electron Microscopic (SEM) photographs and Energy dispersive X-ray spectroscopy (EDS) signature of Bacillus cereus revealed that lead was adsorbed to the cell surface, confirming biosorption capacity of the bacteria [13].
The aim of this study is to identify strains of bacteria isolates from different Pb contaminated sites with high Pb concentrations and to determine the ability of these strains to grow and tolerate high concentrations of Pb.
A number of rhizosphere soil samples were collected from different locations of Saudi Arabia i.e. Al-Riyadh, Al-Taif, Al- Baha where the standing crops were alfalfa, fig and tomato.
Isolation and identification of lead-tolerant bacteria
Ten grams portions from the rhizosphere soil were transferred to a sterile 250 ml sampling bottle containing 90 ml sterilized distilled water. The bottle was shaken manually for 2 min. and farther serial dilutions were made up to 10-5. One ml from each dilution was transferred into sterile petri dishes: nutrient a gar medium containing different concentration of lead 2000, 2500 and 3000 mg/l as PbCl2 was poured into the inoculated petri dishes and mix well. The petri dishes were left for solidification then incubated at 30°C ± 2 for 5 days; the developed colonies were picked up and purified by quadrant streak technique on the same concentration of the respected heavy metal. The purified colonies were examined microscopically to ensure their purity; the pure isolates were kept and periodically sub-cultured on the same heavy metal concentration from which they isolated for the upcoming investigation of the study.
The various isolated microbial strains were identified by Macrogen Company (10F, 254 Beotkkot – ro Geuncheon – qu, Seoul, Rep. of Korea) using 16s ribosomal RNA gene.
The serial dilutions and plate count method using nutrient agar medium supplemented with different concentrations of PbCl2 (2000, 2500 and 3000 mg/L) was used for isolation of Pb tolerant isolates. The growing bacterial colonies on the medium containing the highest concentration of lead (3000 mg/L PbCl2) were picked up and purified then kept on the same nutrient agar medium supplemented with the highest concentration of Pb. The bacterial isolates picked up from the highest concentration of Pb (3000 ppm) were selected for evaluating their abilities in absorbing lead from the liquid medium. Flasks contain sterile 100 ml of nutrient broth medium supplemented with PbCl2 (3000 mg/L) was inoculated with a loop of fresh culture of the respective isolate. The inoculated flasks were incubated at 30°C ± 2 for 7 days in a rotary shaker at 100 rpm. After the incubation period, 25 ml of each culture representing the various isolates were transferred into digestive tubes; a digestive mixture composed of nitric acid and perchloric acid in ratio 1:6 v/v was added to each tube then heated on sand bath till the samples became clear. The digested samples were separately filtered and lead was determined in the filtrate using spectrophotometer (ICP-DES-Oplmia 8000) [14].
Determination of total nitrogen
To differentiate between the various isolated strains, the protein content of various isolates was estimated in their liquid medium. Aliquots of 100 ml of nutrient broth medium supplemented with either lead with concentration 250 mg was distributed into 250 ml sterilized conical flasks each flask was inoculated with 1 ml of freshly prepared inoculum of each isolate then incubated at 30°C ± 2 for 7 days. Total nitrogen in the liquid culture was determined using the semi-microkjeldahl method [15].
Statistical analysis
The obtained results were statistically analyzed using complete randomized block design and program SPSS 17 (2007). ANOVA test was used to differentiate between the various means. Means were compared using LSD at ˂0.05.
From metal industry increase extraction of Cd and Zn from Cd of heavy metals. For example, Bacillus cereus and-rich soil and soil polluted with effluent B. thuringiensis have been shown.
Isolation and efficiency of lead-remediating bacterial strains
Five isolates that showed high tolerance to Pb concentration (3000 mg/L) were obtained from the rhizosphere soil samples that were collected from different locations of Saudi Arabia i.e. Al-Riyadh, Al-Taif, Al-Baha where the standing crops were alfaalfa, fig and tomato, and the identification was carried out by Macrogen Company (10F, 254 Beotkkot–ro Geuncheon–qu, Seoul, Rep. of Korea) using 16s ribosomal RNA gene.
The five bacterial isolates that were identified are two from Al- Baha Serratia marcescens strain WW4 (B1) and Bacillus amyloliquefaciens CC178 (B2), one from Al-Madinah Al- Munawarah B. subtilis strain CCm1999 (M) and two from Al- Taif Bacillus amyloliquefaciens subsp Plantarum Trigo Core 1448 (T1) and Bacillus amyloliquefaciens subsp. Plantarum UCMB 5033 (T2) (Plates 1-5).
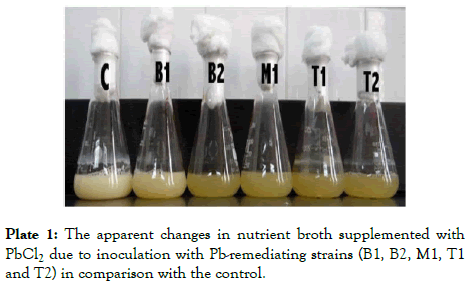

Plate 1: The apparent changes in nutrient broth supplemented with PbCl2 due to inoculation with Pb-remediating strains (B1, B2, M1, T1 and T2) in comparison with the control.
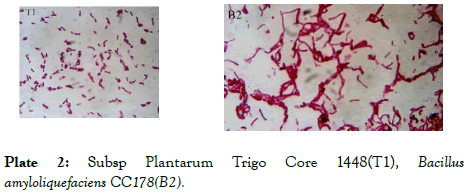
Plate 2: Subsp Plantarum Trigo Core 1448(T1), Bacillus amyloliquefaciens CC178(B2).
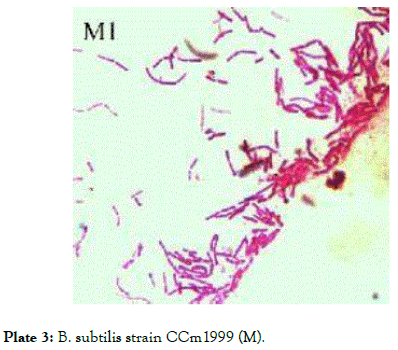
Plate 3: B. subtilis strain CCm1999 (M).
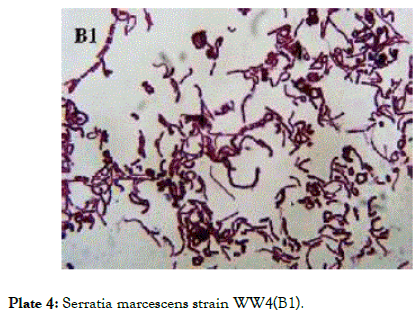
Plate 4: Serratia marcescens strain WW4(B1).
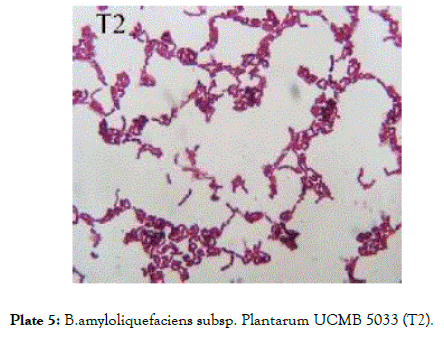
Plate 5: B.amyloliquefaciens subsp. Plantarum UCMB 5033 (T2).
Efficiency of the isolated microbial strains in Pb absorption from the liquid medium
Plate 1 is an overview of the lead-remediating isolates when tested for growth and for their ability to absorb Pb from a 100 ml nutrient broth supplemented with PbCl2 (3000 mg/L) inside 250 ml flasks. The changes in color occurred in the nutrient inside the flasks due to inoculation of the five isolates (T1, T2, M, B1, B2) being clearer in comparison with the non-inoculated medium i.e. the control (C), is an indication of the absorption of Pb by the tested bacterial strains.
Quantitatively, the absorbed amounts of Pb by the five isolated strains, B1, B2, M, T1, T2 were 304, 417, 343, 371 and 189 mg/L respectively (Table 1 and Figure 1), with no significant differences between them except for T2 which gave the lowest Pb content 189 mg/L, while the total amount of Pb measured in the control was 612 mg/L. Bacillus amyloliquefaciens CC178 (B2)., absorbed the highest Pb 417 mg/L, seconded by Bacillus amyloliquefaciens subsp Plantarum Trigo Core 1448 (T1) with 371 mg/L, then B. subtilis strain CCm1999 (M) with 343 mg/L, then Serratia marcescens strain WW4 (B1) with 304 mg/L.The lowest Pb was absorbed by the bacteria Bacillus amyloliquefaciens subsp. Plantarum UCMB 5033 (T2) that reached 189 mg/L.
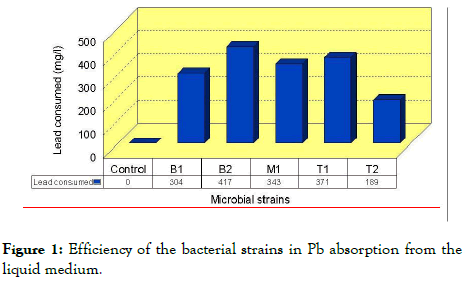
Figure 1. Efficiency of the bacterial strains in Pb absorption from the liquid medium.
| Microbial strains | Amount of lead consumed (mg/l) | Nitrogen content (mg/l) |
|---|---|---|
| Control(612 mg/l) | 00 d | 2b |
| B1 | 304b | 7a |
| B2 | 417a | 8a |
| M1 | 343ab | 7a |
| T1 | 371ab | 8a |
| T2 | 189c | 7a |
| LSD | 109 | 1.12 |
Table 1: Efficiency of the isolated microbial strains in lead absorption and nitrogen produced in the liquid medium.
In addition the amount of N content in the liquid cultures of the various bacterial strains identified and the control treatment was assessed (Table 1) and ranged between 7 and 8 mg/L being the highest with Bacillus amyloliquefaciens CC178 (B2) and amyloliquefaciens subsp Plantarum Trigo Core 1448 (T1) but significantly high compared with the control which gave only 2 mg/L.
B1 and B2 are the strains isolated from Al-Baha governorate; M1 is the strain isolated from AlMadinah governorate; T1 and T2 are the strains isolated from Al-Taif governorate.
Contamination of heavy metals in the environment is a major global problem because they are toxic and create many health problems to human and animal and aquatic life specially Pb contamination. The first step in bioremediation strategy is to identify candidate bacteria strains capable of absorbing Pb from contaminated sites. The test of the ability of the five isolated bacteria strains to absorb Pb from liquid medium supplemented by PbCl2 (3000 mg/L) revealed that four of the isolates, Bacillus amyloliquefaciens CC178 (B2), absorbed the highest Pb percentage 68.16%, seconded by Bacillus amyloliquefaciens subsp Plantarum Trigo Core 1448 (T1) with 60.62% Pb, then B. subtilis strain CCm1999 (M) with Pb percentage of 56.60%, then Serratia marcescens strain WW4 (B1) with 49.67%. The lowest Pb percentage absorbed was by the bacteria Bacillus amyloliquefaciens subsp. Plantarum UCMB 5033 (T2) that reached 30.88% compared to the control with 612 mg/L. The varying responses of the strains to Pb concentrations might be due to variation in resistance mechanisms.
This finding is in agreement with Mohammad et al. [16] who found that within 30 minutes Bacillus thuringiensis strain OSM29 can remove Pb from Ph contaminated soil at a biosorption capacity of 90% [13]. Also identified six Pb resistant bacterial strains one of them is Bacillus cereus, and [17] in Iran isolated a bacteria of the genus Halomonas that can uptake more than 90% Pb [12]. Reported that the role played by soil microbes in Pb bioremediation is either through absorption, transformation or degradation [18,19]. Used the bacteria Pseudomonas sp., Chryseomonas lutesla and Bacillus circulans to remove Pb from industrial wastes. Absorption of Pb from the nutrient broth amended with 3000 mg/L Pb by the isolated bacteria is most probably takes place through adsorption to the cell surface, as this was confirmed by Murthy et al. [13], who used scanning electron microscopic (SEM) photographs and Energy dispersive X-ray spectroscopy (EDS) and signature of Bacillus cereus revealed that lead was adsorbed to the cell surface, confirming biosorption capacity of the bacteria. Also Pb absorption by bacteria may be through other different mechanisms, adsorption/desorption as said by Mamaril et al. [20] or through pumping of Pb ions exterior to the cell, and accumulation and sequestration of Pb ions inside the cell, or biotransformation of the toxic form of Pb to less toxic form as said by Thacker et al. [21].
Five Pb-tolerant bacteria strains were isolated and identified, two were isolated from the root rhizosphere soil cultivated with alfalfa plants at Al-Baha Serratia marcescens strain WW4 (B1) and Bacillus amyloliquefaciens CC178 (B2), one from the rhizosphere soil under fig plants at Al-Madinah Al-Munawarah B. subtilis strain CCm1999 (M) and two from the rhizosphere soil under tomato plants at Al-Taif Bacillus amyloliquefaciens subsp Plantarum Trigo Core 1448 (T1) and Bacillus amyloliquefaciens subsp. Plantarum UCMB 5033 (T2). Four of the isolates showed high Pb absorption percentages Bacillus amyloliquefaciens CC178 (B2), absorbed the highest Pb percentage 68.16%, seconded by Bacillus amyloliquefaciens subsp Plantarum Trigo Core 1448 (T1) with 60.62% Pb, then B. subtilis strain CCm1999 (M) with Pb percentage of 56.60%, then Serratia marcescens strain WW4 (B1) with 49.67%. The lowest Pb percentage absorbed was by the bacteria Bacillus amyloliquefaciens subsp. Plantarum UCMB 5033 (T2) that reached 30.88% compared to the control with 612 mg/L. So it can be said that all of these isolated Pb resistant strains are well efficient for bioremediation of Pb from contaminated sites.
My thanks are to my department colleagues and technicians for their continuous support, encouragement and providing me with all the required facilities to achieve the practical work of the thesis, and also to my family for their constant support.
Citation: Al-Hazmi RHM (2019) Isolation and Identification of Five Lead Bio-Remediating Bacterial Strains. J Plant Biochem Physiol. 7:241. doi: 10.35248/2329-9029.19.7.241
Received: 15-Jul-2019 Accepted: 12-Aug-2019 Published: 19-Oct-2019
Copyright: © 2019 Al-Hazmi RHM, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.